 CLART CMA is an in vitro diagnostics test line of products for the detection of mutations in genes associated with response to therapy in cancer patients. Products CLART CMA are based on CLART Technology, and in particular on Multiplex PCR amplification followed by visualization in low-density arrays.
CLART CMA is an in vitro diagnostics test line of products for the detection of mutations in genes associated with response to therapy in cancer patients. Products CLART CMA are based on CLART Technology, and in particular on Multiplex PCR amplification followed by visualization in low-density arrays.
CLART CMA KRAS·BRAF·PI3K:
Detection and differentiation of 15 point mutations associated to response to therapy in patients with metastatic colorectal cancer.
CLART CMA NRAS·iKRAS:
Detection and differentiation of 10 point mutations associated to response to therapy in patients with metastatic colorectal cancer.
The exact mutations to be detected with CLART CMA KRAS·BRAF·PI3K and CLART CMA NRAS·iKRAS kits are:
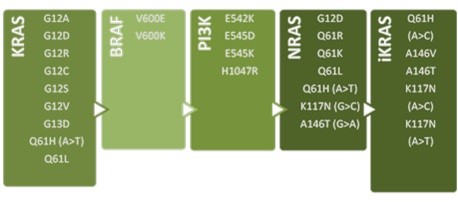
The design of these kits allows the user to perform an analysis on demand by combining the different amplification references according to the necessities of the lab, and visualizing all together using just one array per kit.
CLART CMA EGFR:
Detection and differentiation of 40 point mutations, deletions and insertions associated with response to therapy in non-small-cell lung cancer patients. The exact mutations to be detected with CLART CMA EGFR kit are:
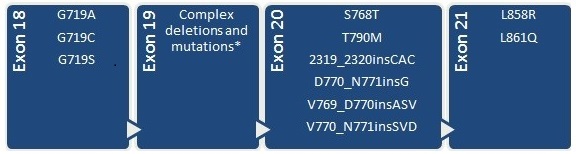
CLART CMA ALK · ROS1
Detection and genetic identification of major chromosomal translocations in ALK and ROS1 genes associated with response to therapy in patients with lung cancer.
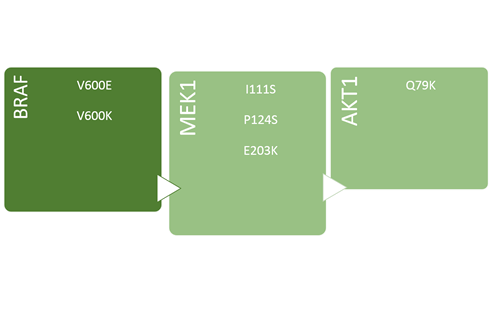
CLART CMA BRAF·MEK1·AKT1:
Detection and differentiation 6 point mutations associated with response to therapy in patients with melanoma.
Advantages of CLART CMA diagnostics:
Minimum amounts of sample are required, the remaining tissue being preserved for future or parallel studies.
All reagents and components are supplied ready to use, the hands-on time invested on the technique being significantly reduced.
CLART CMA diagnostics constitutes an optimization of existing protocols, improving the laboratory workflow.
It is a tool to detect chimeric tumors, to distinguish between primary and metastatic tumors, to establish the most adequate treatment or to predict response to medication.
